The preprint describing MoDLE's capabilities, design and implementation is available on bioRxiv at 10.1101/2022.04.13.488157.
MoDLE is released under a permissive open-source license, and its source code is available on GitHub.
Paper abstract and fig. 1
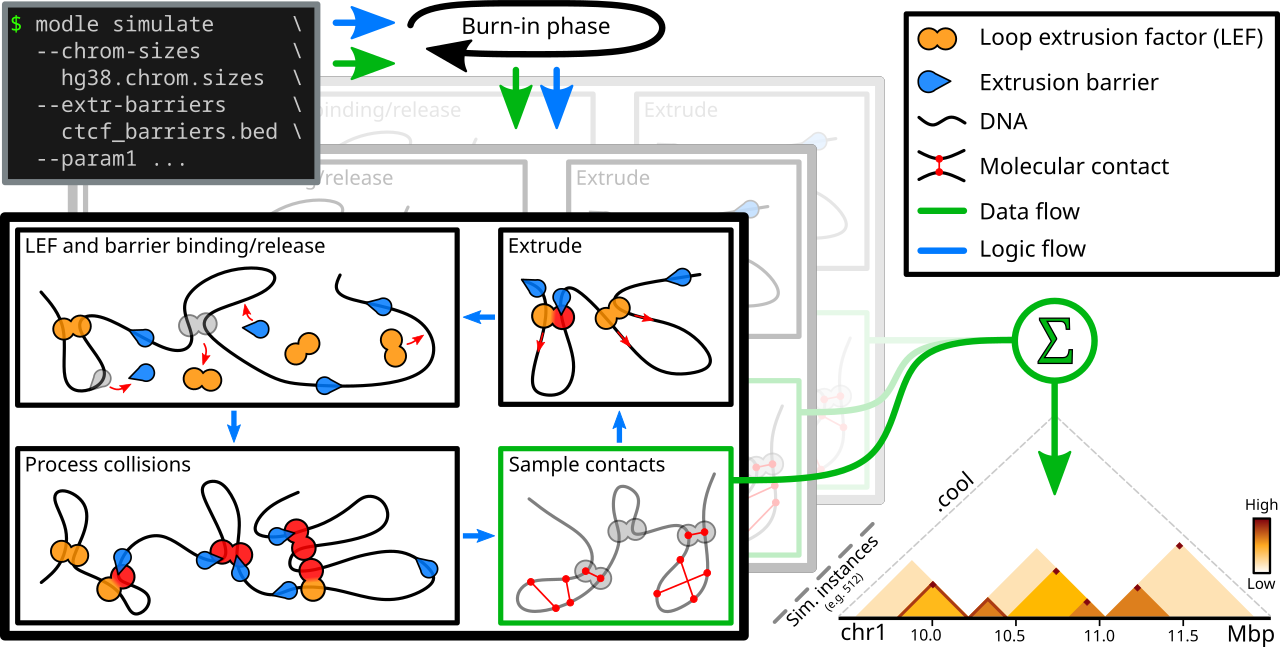
Schematic and simplified overview of MoDLE. Input files specify genome regions to be simulated (e.g. a chrom.sizes file) and their barrier positions (e.g. CTCF binding sites and orientation) in BED format. Optional parameters control the specifics of a simulation. Loop Extruding Factors (LEFs) bind to, extrude and release from the regions and interact with modeled barriers according to input parameters. Loop extrusion and intra-TAD contacts of a randomized subset of loops are recorded each epoch and aggregated into an output cooler file containing the final simulated contact frequencies. Simulation halts when a target number of epochs or a target number of loop extrusion contacts have been simulated.